About Marina Sapir, Ph.D.
Resume
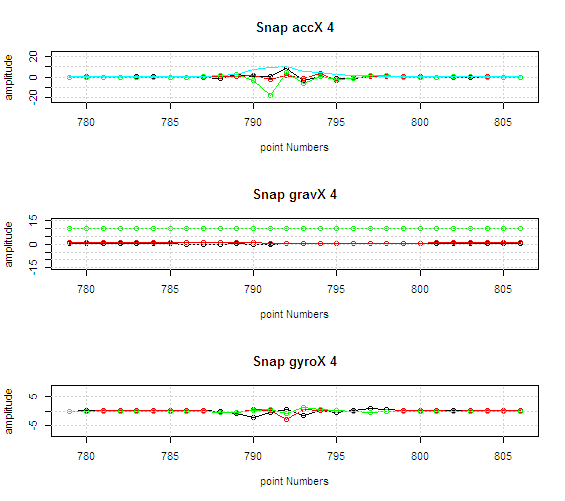
 Data researcher with wide expertise in machine learning, data analysis, signal processing
Data researcher with wide expertise in machine learning, data analysis, signal processing
Solving customer's machine learning and data analysis problems from economics, finance, ecology, signal processing and other areas of study.
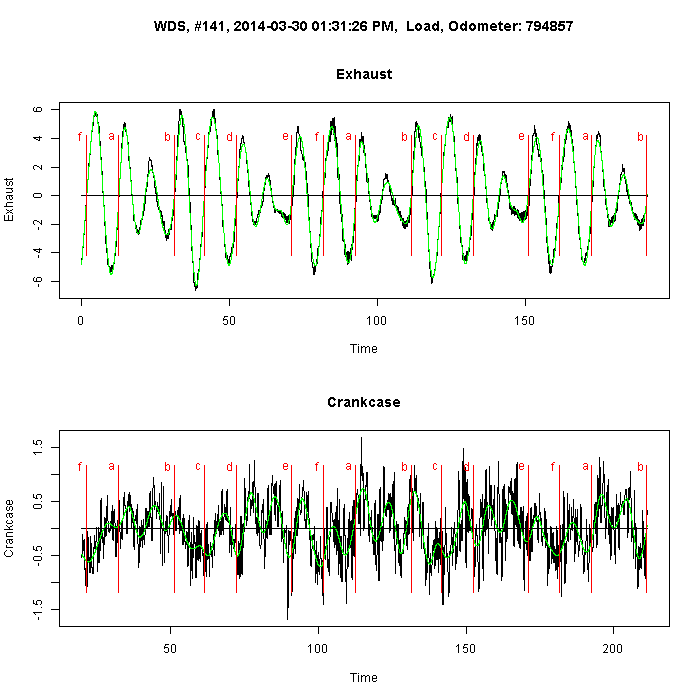
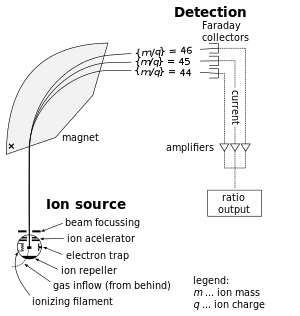
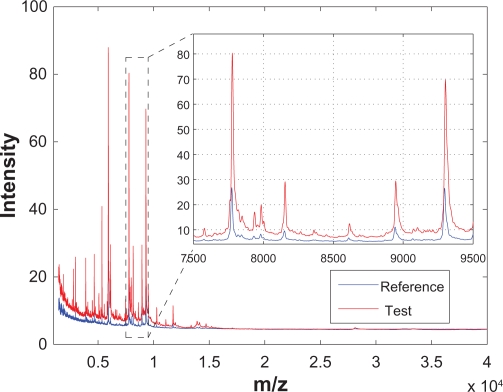
Analysis of mass spectrometry data for medical diagnostics. New algorithms for spectrometry data processing. Machine learning with small datasets and large number of features.
Medical image processing, development of methods and aggregated predictive features for survival analysis with medical applications.
Developed methods and software for computerized drug discovery. Creating commercial application with C++.
Assessment of differential gene expression for RNA microarrays based on a single array. The idea served as foundation of many further statistical studies in various institutions, including University of Berkeley. Designed and developed GUI software for modeling of microarray data. Presented the results on the conference and made invited talks on this subject in universities and commercial companies.
-
Ph.D., Computer Science. Dissertation: Discovery of optimal logical rules in data. Institute of Automated Control, Russian Academy of Science, Moscow, Russia
Master of Science., Mathematics, Ural State University, Russia